Overview
In the previous post I briefly touched on DICOM as the crucial standard in medical imaging for both data exchanging and data storage. It is important to understand that DICOM is such a massive standard that, beyond data exchanging and storage, has expanded into many different areas around imaging, that no device (or information system) can ever implement every single aspect of the standard. A device or information system complies to (and implements) a subset of the DICOM standard. The manufacturer must provide a document (DICOM conformance statement) to spec out which parts of the standard are implemented. Care providers (e.g. hospitals) are supposed to review these specs as part of the procurement process to ensure interoperability with existing information system.
Medical Imaging Informatics
When it comes to diagnosis, historical examinations provide baseline reference for radiologist. They are sometimes even more revealing than current imaging data acquired from a patient. Compared to the current exams, historical ones are referred to as priors. Priors are useful only if they are relevant to the current exams in terms of body part, modality, and exam procedure (the particular problem being studied). The effort to find out and pre-load relevant priors so they are ready to display along with current exams, is called “prefetch”. With huge demand in exchanging imaging data, “prefetch” has developed into its own sub-market in imaging informatics industry.
Trust me, this is a difficult undertaking (and why I was in the industry). First, patient identities in each healthcare organization are usually different for lack of universal medical record number; patient’s name can change (marriage, divorce, or just for fun); or it can be unavailable, if patient is simply not in a condition to provide identify (e.g. trauma). Second, even if you get “who’s who” right, you’d have to dig into all his history for useful information from several different systems. The definition of relevant prior can be different depending on the specific medical specialty. Then, the old data are typically stored in a slow part of the storage from their source system, yet the patient might be bleeding and dying on the table, waiting for prior retrieval like pulling teeth. Last, but not least, the priors being retrieve might be from a modality of previous generation from 1990s; good luck with current display application. Sorry that sounds a lot but in real life, there are even more challenges. In General, there are four categories of applications that need to support DICOM:
- Acquisition devices: Modalities need to store newly acquired studies persistently;
- Routing applications: usually by the name of some routers, gateways or bridges that receive imaging studies, decorate the metadata (because a lot of legacy devices can’t do it), and send to one or multiple defined destination;
- Archives (e.g. PACS, VNA): They usually use dedicated database (metadata) and storage systems (pixel data). They are the repository of imaging data and must provide full support of transfer capability;
- Peripheral applications that uses DICOM data, such as 3D post-processing or DICOM testing (grassroot dicom, dcmtk)
No matter what the devices are, they must follow certain protocols in order to communicate with each other.
Data structure and encoding
DICOM PS3.5 defines data types in VR (value representation). Each VR has its own purpose, allowed characters, and length limit. For example:
| VR | Definition | Allowed Characters | Length Limit |
|---|---|---|---|
| SH Short String | A string of characters | 16 maximum | |
| LO Long String | A string of characters | 64 maximum | |
| AE Application Entity | A string of characters that identifies a DICOM application running on a compliant device | 16 maximum | |
| CS Code String | A string to represent code | 16 maximum | |
| PN Person Name | Person’s name, with caret (^) as delimiter | 64 maximum | |
| UI Unique Identifier | An ID that uniquely identify an item, such as 1.2.840.100008.1.1 | 0-9 and period (.) | 64 maximum |
| DA Date | A string to represent date YYYYMMDD | 0-9 | 8 |
| US Unsigned Short | Unsigned binary integer, 16 bits long | 2 | |
| SQ Sequence of other items | Sequence of other items | ||
| UN Unknown | A string of bytes where the encoding of contents is unknown |
DICOM metadata is a dataset comprised of a set of data elements. Each element includes tag, (optional) VR, length of value, and the actual value, as shown below:
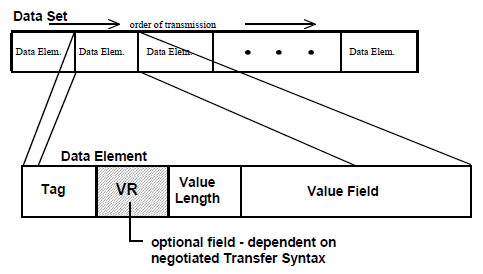
VR is optional because it can be implicitly determined based on DICOM data dictionary defined in PS3.6. The most common tags are:
| Level | Tag | Meaning | VR |
|---|---|---|---|
| Patient | 0010,0010 | Patient’s Name | PN |
| Patient | 0010,0020 | Patient ID | LO |
| Patient | 0010,0021 | Issuer of Patient ID | LO |
| Patient | 0010,0024 | Issuer of Patient ID Qualifier Sequence | SQ |
| Patient | 0010,0030 | Patient’s Birth Date | DA |
| Study | 0008,0020 | Study Date | DA |
| Study | 0008,0050 | Accession Number | SH |
| Study | 0008,0061 | Modalities In Study | CS |
| Study | 0008,1030 | Study Description | LO |
| Study | 0020,000D | Study Instance UID | UI |
| Study | 0020,0010 | Study ID | SH |
| Series | 0008,103E | Series Description | LO |
| Series | 0008,0015 | Body Part Examined | CS |
| Series | 0008,0060 | Modality | CS |
| Series | 0020,0011 | Series Number | IS |
| Series | 0020,000E | Series Instance UID | UI |
| Series | 0020,0060 | Laterality | CS |
| SOP inst | 0008,0016 | SOP Class UID | UI |
| SOP inst | 0008,0018 | SOP Instance UID | UI |
| SOP inst | 0012,0010 | Transfer Syntax UID | UI |
| SOP inst | 0020,0013 | Instance Number | IS |
When VR is not explicitly spelled out, the data encoding is known as implicit VR. The opposite is explicit VR, where each data element spells out the VR type. When storing numeric value, such as tags or numeric values for tags, the predominant format stores lower byte before higher bytes, known as little endian. Rarely seen in DICOM is big endian, the opposite order of storing numeric values.
The tag include a group number (e.g. 0020) and element number (e.g. 0013). If group number is odd, it is a private tag not defined in PS3.6
The actual length of tag value shall always be even number. Odd-sized value should add an additional character (e.g. trailing space), known as even-length padding to meet this requirement.
Above is the basic rules for DICOM data structure and encoding. For more information about encoding, including transfer syntax, refer to this previous post.
Transactions (DIMSEs and SOP classes)
DICOM has its own information model of real world. It requires some clinical knowledge to come to full understanding. For technical people, we just need to understand the patient-study-series-image hierarchy.
- One patient may have multiple studies
- Each study may include one or more image series
- Each series has one or more images
Images are also referred to as SOP instance, a general terms that include not only images, but also reports, and other types of objects.
A business transaction in DICOM is termed DIMSE, for example:
| Name | Group | Type | Description |
|---|---|---|---|
| C-STORE | DIMSE-C | Operation | A stores an image to B |
| C-MOVE | DIMSE-C | Operation | A tells B to store an image to C |
| C-FIND | DIMSE-C | Operation | Query for patient, study, series, images |
| C-ECHO | DIMSE-C | Operation | DICOM level “ping” |
| N-EVENT-REPORT | DIMSE-N | Notification | Report an event |
| N-ACTION | DIMSE-N | Operation |
Some DIMSEs involves multiple SOP classes based on the type of image being processed. For example, SOP Class UID (10.2.840.10008.5.1.4.1.1.1) represents storage for CR image. A c-store transaction includes:
- Requestor (C-STORE SCU) sends a request (C-STORE-RQ), to store specified SOP class in certain transfer syntax
- The request is followed by the actual data (PDU)
- Once completed, the Response (C-STORE SCP) respond with C-STORE-RP
Handshake (Association)
In the previous C-STORE example, proper syntax that are mutually supported must be used in order for the requestor and receiver to process the transaction. Both parties learn each other’s supported transfer syntax through an upfront handshake process known as DICOM association.
In DICOM association, the initiating party presents a list of supported pairs of SOP class and transfer syntax. Each pair is called a presentation context, and the responding party must respond to each presentation context in the proposed list, with either yes or no.
This negotiation is similar to the cipher negotiation in TLS handshake.
For more detailed information, please refer to Pianykh’s book “DICOM, a practical introduction and survival guide“.
